Introduction
A biological network is a method of representing systems as complex sets of binary interactions or relations between various biological entities. Therefore, graphs can be used to capture interactions between molecules such as DNA; RNA, proteins or metabolites.
A typical graphing representation is made of a set of nodes connected by edges.
This resource will focus on the history of the graph theory and will describe the multiple biological networks that exist.
History & Description
The graph theory emerged in 1736 when Leonhard Euler analyzed a real-world issue known as the Seven Bridges of Königsberg (1) a mathematical problem that Euler proved to be impossible to solve.
Later on, between the 1930’s and 1950’s; random graph were theorized. It describes a probability distributions over graphs, lying at the intersection of graph theory and probability theory. Afterward, in the 90’s, it was discovered that many different types of “real” networks had structural properties quite different from random networks.
Finally, the late 2000’s began shaping the emergence of systems biology, network biology, and network medicine (2). In 2014, graph theoretical methods were used to analyze biological networks (3).
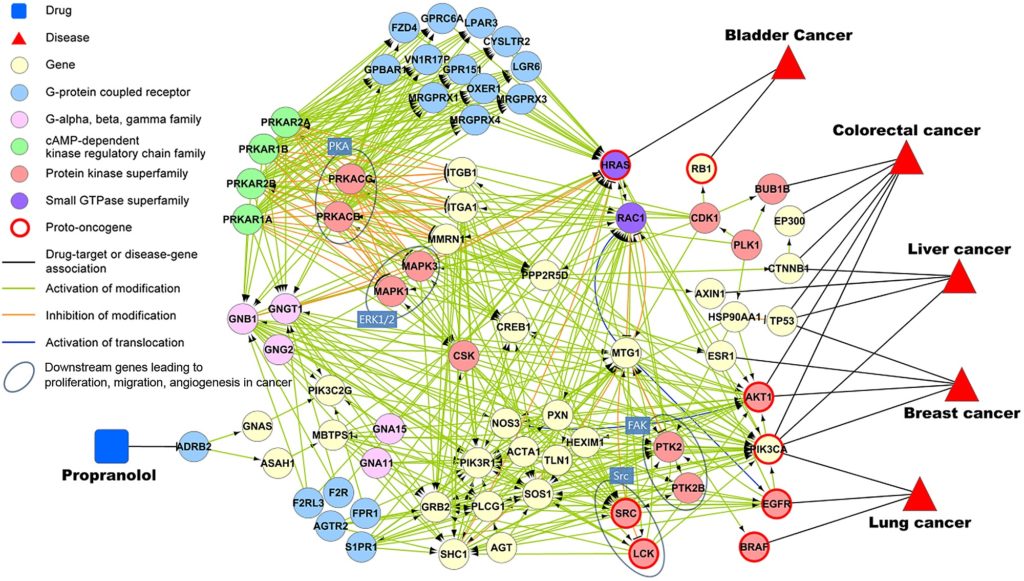
Biological Networks
Multiple types of networks exist in Biology:
- Protein-protein interaction networks represents the physical relationship among proteins (depicted as nodes). Their interactions are represented as undirected edges. Protein-protein interactions (PPI) are traditionally discovered using experimental techniques like yeast two-hybrid system, or more recently high-throughput studies using mass spectrometry. Many international efforts have resulted in databases that catalog experimentally determined PPIs (like MINT, IntAct, BioGRID, etc…) or computationally predicted PPIs (Examples : FunCoup, STRING).
- Gene regulatory networks (DNA-protein interaction networks), also called GRN, are represented with genes and transcriptional factors as node, and the relationship between them as directional edges (either promoting gene regulation or its inhibition). GRNs are usually constructed using available databases and the knowledge therein, such as Reactome and KEGG. In order to discover these interactions, high-throughput measurements, such as microarray, RNA-seq, ChIP-seq, etc., are used to gather large-scale transcriptomics data.
- Signaling networks comes from the signal transduction within cells. These signals play essential role in the tissue structure. These networks usually incorporate PPI, GRN and metabolic networks.
- Other type of biological networks exist. Here a non-exhaustive lists that we won’t describe further in this article:
- Gene co-expression networks
- DNA-DNA chromatin networks
- Metabolic networks
- Neuronal networks
- Food webs
- Between-species interaction networks
- Within-species interaction networks
2.2.Modelling biological networks
In order to get useful information from a biological network, an understanding of the statistical and mathematical techniques of identifying relationships within networks is vital.
To provide insight into the relationships of a networks it is essential to identify « Association », « Communities » and « Centrality ».

Association is the type of measures used for nodes relationships in order to analyze a network. In biological networks, biologists would use Correlation in order to measure to analyze a network.
Centrality is a concept that can be extremely useful to analyze biological network structures and therefore to measure the prominence of a node in a network. Multiple ways of measuring centrality exist: betweeness, degree, Eigenvector, Katz centrality. Even though each type can provide different insights on nodes, they all measure the prominence of a node.
Communities correspond to the network subdivision into groups of nodes representing like-regions. By doing so, it provides a better view of the pockets of highly connected relationships. As of today, scientists continue to invent new ways of sub sectioning networks and therefore algorithms to create these relationships. From these algorithms, two are very commonly used in biological networks for community detections: The Louvain Method (4) and the Leiden Algorithm.
- The Louvain Method attempts to maximize modularity, which favors heavy edges within communities and sparse edges between, within a set of nodes. There is some limits on the Louvain method : it could create badly connected communities by degrading a model for the sake of maximizing a modularity metric.
- The Leiden Algorithm adds up on the Louvain Method by providing a number of improvements. Among them, when joining nodes to a community, only neighborhoods that have been recently changed are considered (this will increase sharply the speed of merging nodes). Also, Leiden randomly chooses for a node from a set of communities to merge with; allowing for greater depth in choosing communities (Louvain solely focuses on maximizing the modularity that was chosen). The Leiden algorithm, while more complex, performs faster with better community detection (5).
References
- https://en.wikipedia.org/wiki/Seven_Bridges_of_K%C3%B6nigsberg
- Koutrouli, M., Karatzas, E., Paez-Espino, D. & Pavlopoulos, G.A. A Guide to Conquer the Biological Network Era Using Graph Theory. Front. Bioeng. Biotechnol. 8:34 (2020). doi: 10.3389/fbioe.2020.00034
- Emmert-Streib, F., Dehmer, M. & Haibe-Kains, B. Gene regulatory networks and their applications: understanding biological and medical problems in terms of networks. Front. Cell Dev. Biol. 2:38 (2014). doi: 10.3389/fcell.2014.00038
- Blondel, V.D., Guillaume, J.L., Lambiotte, R. & Lefebvre, E. Fast unfolding of communities in large networks. J. Stat. Mech. (2008). doi: 10.1088/1742-5468/2008/10/P10008
- Traag, V.A., Waltman, L. & van Eck, N.J. From Louvain to Leiden: guaranteeing well-connected communities. Sci Rep9, 5233 (2019). https://doi.org/10.1038/s41598-019-41695-z



