Introduction
Methods for analyzing single-cell data perform a core set of computational tasks, ranking from dimensionality reduction, cell clustering, removal of unwanted variation, cell-state annotation, identification of spatial patterns of gene expression, etc. Most of these methods rely on probabilistic models that provide principled ways to capture uncertainty in biological systems.
Despite the appeal of these models, multiple problems hamper their wide adoption.
In this article we will describe the Python library scVI and how it tries to circumvent these obstacles.
History & Challenges of scVI
ScVI originated in 2017, in Yosef Lab, located at UC Berkeley (1). Its development was centered around reducing the barrier to entry for using probabilistic models in single-cell omics analyses.
As described previously, probabilistic models have several problems to resolve. First, it can be difficult to implement and run for the end user. This is often due to the requirement of interacting with Python-based machine-learning libraries. Second, working with higher-level machine-learning packages, like the popular PyTorch or Keras, does not work seamlessly with single-cell omics data.
ScVI was created to resolve these two issues and bridge together the world of deep learning models and single-cell data analysis.
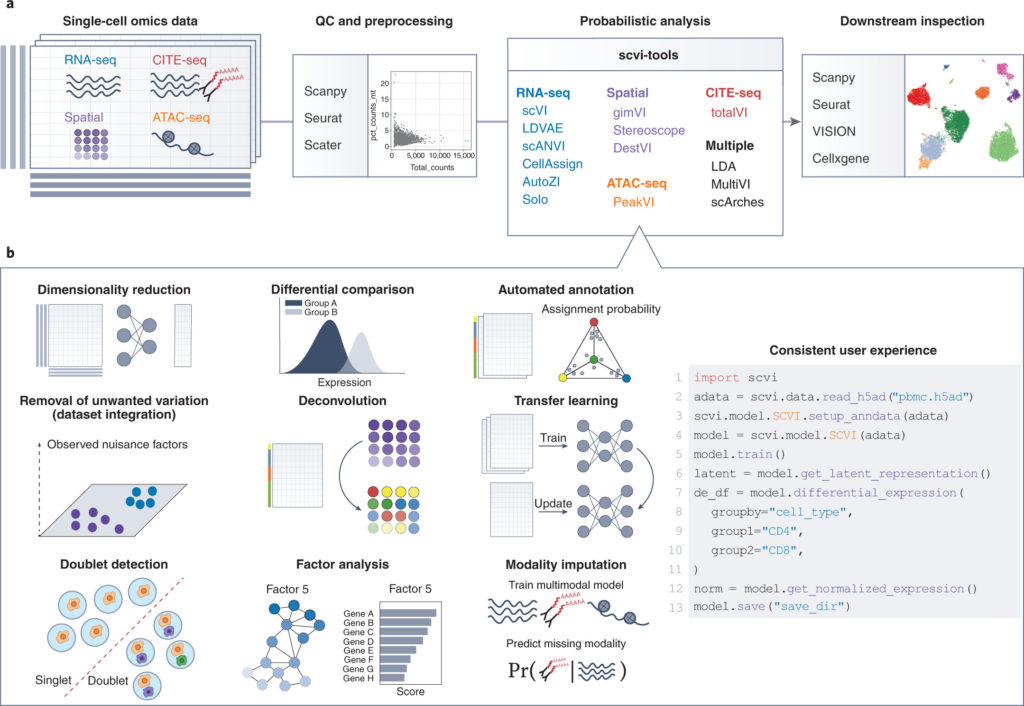
Figure 1. User perspective of scvi-tools. a. Overview of single-cell omics analysis pipeline with scvi-tools. b. Overview of the functionality of the models implemented in scvi-tools, with a simple and consistent user interface.
From the end user’s perspective sci-tools offers standardized access to a variety of methods to tackle single-cell data analysis tasks, from scRNA-seq data, annotation of single-cell profiles, deconvolution of bulk spatial transcriptomics profiles and multi-modal analysis of CITE-seq data. All 14 models currently implemented in scvi-tools interact with ScanPy through the annotated dataset (AnnData (2, 3)) format and the models share a consistent user interface.
In order to comply with the second obstacles, regarding deep learning models integration, scvi-tools offers set of building blocks that make it easy to implement or modify new or existing models. Thus, it can implement libraries such as PyTorch (4), PyTorch Lightning (5, 6) and Pyro (7, 8) and facilitate probabilistic models design with neural network components and GPU acceleration.
The deployment and continuous development of scvi-tools gives developers the opportunity to adhere to standard API and coding conventions; giving more accessibility for new users. Hence the single-cell community at large will be better serve to prototype new models and enhancing the scientific discovery pipeline.
References
- https://yoseflab.github.io/software/scvi-tools/
- Virshup, I., Rybakov, S., Theis, F.J., Angerer; P., Wolf, F.A. anndata: Annotated data (2021). doi:10.1101/2021.12.16.473007
- https://github.com/scverse/anndata
- Paszke, A., Gross, S., Massa, F., Lerer, A., Bradbury, J., Chanan, G., et al. PyTorch: An Imperative Style, High-Performance Deep Learning Library. NeurIPS. 32 (2019).
- The lightweight PyTorch wrapper for ML researchers
- https://github.com/Lightning-AI/lightning
- https://github.com/pyro-ppl/pyro
- Bingham, E., Chen, J.P., Jankowiak, M., Obermeyer, F., Pradhan, N., Karaletsos, T. Pyro: Deep Universal Probabilistic Programming 20(28):1−6 (2019).



